This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is Protein Homology?
Protein homology is a measure of evolutionary relationships between proteins [1]. Homologous proteins have a significant amount of sequence similiarity and usually have similar domains [2]. HomoloGene and BLAST are online tools from NCBI used to find protein or DNA homologs or measure sequence similarity.The maximum identity numbers from BLAST listed below each protein show what percent of the protein matches when exactly lined up with the Human ANK1, however the proteins may actually be much more similar due to insertions or deletions changing the frame of alignment. Sequence alignments can be seen below.
ANK1 Homologous Proteins
|
Human - Homo sapiens ANK1
Accession Number: NP_000028.3 FASTA Chimpanzee - Pan troglodytes ANK1 Accession Number: XP_003311743 FASTA Max Identity: 99% Rhesus Macaque - Macaca mulatta ANK1 Accession Number: XP_001099591 FASTA Max Identity: 95% Dog - Canis lupus familiaris ANK1 Accession Number: XP_539957 FASTA Max Identity: 89% Cow - Bos taurus ANK1 Accession Number: DAA14471 FASTA Max Identity: 90% Mouse - Mus musculus Ank1 Accession Number: NP_001104253 FASTA Max Identity: 91% Nematode Worm- Caenorhabditis elegans Unc-44, isoform f Accession Number: NP_001021268 FASTA Max Identity: 54% |
Brown Rat - Rattus norvegicus Ank1
Accession Number: NP_001100792 FASTA Max Identity: 75% Chicken - Gallus gallus ANK1 Accession Number: XP_424401 FASTA Max Identity: 82% Zebrafish - Danio rerio Predicted ankyrin-1-like Accession Number: XP_003200911 FASTA Max Identity:67% Arabidopsis - Arabidopsis thaliana ATSG14230 Accession Number: NP_196927 FASTA Max Identity: 27% Asian Rice - Oryza sativa Os02g0457500 Accession Number: NP_001046778 FASTA Max Identity: 27% Budding Yeast - Saccharomyces cerevisiae Akr1p Accession Number: EDV08055 FASTA Max Identity: 33% Fruit Fly- Drosophila melanogaster Ank2-PJ Accession Number: NP_001097538 FASTA Max Identity: 51% |
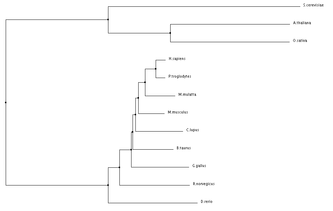
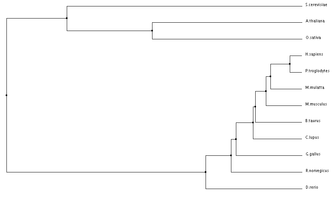
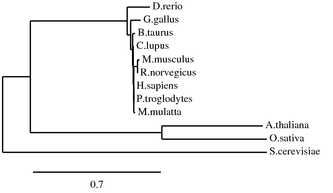
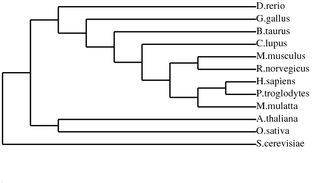
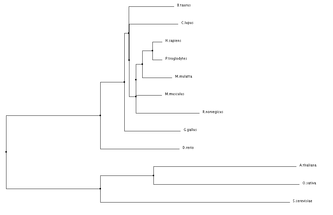
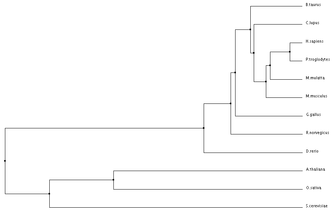
ANK1 Homolog Alignments and Phylogenetic Trees
Programs such as the ones used below can be used to align protein sequences and create phylogenetic trees. Phylogenetic trees are a visual representation of evolutionary relationships, and in this case are based on the protein sequence similarities. Click on Images for full size.
Clustal Omega
|
Phylogeny.fr |
T-Coffee |
||||||
Analysis
Since ANK1 is an erythrocytic (red blood cell) protein, it's unsurprising to see that many animals have proteins similar to it. What is more interesting is that there are proteins similar to human ANK1 found in arabidopsis (a plant) and yeast (a fungus). Just because the sequences are similar does not necessarily mean that the protein activity is also similar, but if it is both organisms could serve as useful model organisms for further research into the mutations that cause spherocytosis.
References
1. Reeck, G.R, de Haën C, Teller D.C, Doolittle R.F, Fitch W.M, Dickerson R.E, Chambon P, McLachlan A.D, Margoliash E, Jukes T.H, Zuckerkandl E, “Homology” in proteins and nucleic acids: A terminology muddle and a way out of it, Cell, Volume 50, Issue 5, 28 August 1987, Page 667, ISSN 0092-8674, 10.1016/0092-8674(87)90322-9. Retrieved from: http://www.sciencedirect.com/science/article/pii/0092867487903229
2. Garrett, R.H., Grisham, C.M. 1999. Biochemistry Second Edition Saunders College Publishing. p.142
3. Geer LY, Marchler-Bauer A, Geer RC, Han L, He J, He S, Liu C, Shi W, Bryant SH. The NCBI BioSystems database. Nucleic Acids Res. 2010 Jan; 38(Database issue):D492-6.(Epub 2009 Oct 23) [PubMed PMID: 19854944]
4. Altschul S.F., Gish W., Miller W., Myers E.W. and Lipman D.J. (1990) Basic local alignment search tool. J. Mol. Biol. 215: 403-410.
5. Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins D Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega Molecular Systems Biology 7 Article number: 539 doi:10.1038/msb.2011.75
6. Goujon M, McWilliam H, Li W, Valentin F, Squizzato S, Paern J, Lopez R (2010) A new bioinformatics analysis tools framework at EMBL-EBI Nucleic acids research 2010 Jul, 38 Suppl: W695-9 doi:10.1093/nar/gkq313
7. Henikoff J.G. Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci U S A. 1992 November 15; 89(22): 10915–10919. PMCID: PMC50453
8. Eddy, S.R. Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology 22, 1035 - 1036 (2004) doi:10.1038/nbt0804-1035
9. Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. (PubMed)
10. Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9. Epub 2008 Apr 19. (PubMed) *: joint first authors
11. Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, Mar 19;32(5):1792-7. (PubMed)
12. Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000, Apr;17(4):540-52. (PubMed)
13. Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, Oct;52(5):696-704. (PubMed)
14. Anisimova M., Gascuel O. Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 2006, Aug;55(4):539-52. (PubMed)
15. Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 2006, Oct 10;7:439. (PubMed)
2. Garrett, R.H., Grisham, C.M. 1999. Biochemistry Second Edition Saunders College Publishing. p.142
3. Geer LY, Marchler-Bauer A, Geer RC, Han L, He J, He S, Liu C, Shi W, Bryant SH. The NCBI BioSystems database. Nucleic Acids Res. 2010 Jan; 38(Database issue):D492-6.(Epub 2009 Oct 23) [PubMed PMID: 19854944]
4. Altschul S.F., Gish W., Miller W., Myers E.W. and Lipman D.J. (1990) Basic local alignment search tool. J. Mol. Biol. 215: 403-410.
5. Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins D Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega Molecular Systems Biology 7 Article number: 539 doi:10.1038/msb.2011.75
6. Goujon M, McWilliam H, Li W, Valentin F, Squizzato S, Paern J, Lopez R (2010) A new bioinformatics analysis tools framework at EMBL-EBI Nucleic acids research 2010 Jul, 38 Suppl: W695-9 doi:10.1093/nar/gkq313
7. Henikoff J.G. Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci U S A. 1992 November 15; 89(22): 10915–10919. PMCID: PMC50453
8. Eddy, S.R. Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology 22, 1035 - 1036 (2004) doi:10.1038/nbt0804-1035
9. Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. (PubMed)
10. Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9. Epub 2008 Apr 19. (PubMed) *: joint first authors
11. Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, Mar 19;32(5):1792-7. (PubMed)
12. Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000, Apr;17(4):540-52. (PubMed)
13. Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, Oct;52(5):696-704. (PubMed)
14. Anisimova M., Gascuel O. Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 2006, Aug;55(4):539-52. (PubMed)
15. Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 2006, Oct 10;7:439. (PubMed)